Projects & Research Interests
Although my research interests and projects are presented here from an individual point of view ("homepage"), all results are obviously a collective effort. See our publications for details about colleagues and collaborators.
Software technologies to foster image-based scientific collaboration and open science

In 2010, I initiated the CYTOMINE research project which lead to the development of the CYTOMINE open-source web software platform (Marée et al., Bioinformatics 2016; Rubens et al., 2019). Cytomine is a "Google/OpenStreet Maps"-like rich internet application for remote visualization, collaborative annotation and automated analysis of high-resolution, multi-gigapixels images.
It is now actively used in various domains (including digital pathology, large-scale microscopy, and other fields beyond the biomedical field) by various entities around the world collaborating over the web.
We are continuously developing new software modules and algorithms for multimodal data sources (I'm was co-lead of the Software workpackage of the NEUBIAS and COMULIS COST ACTIONs), benchmarking, and applications in various fields (see CYTOMINE research project page). In addition to our ongoing research & development at University of Liège, I'm co-founder of Cytomine SCES, a not-for-profit cooperative company established to coordinate the open-source community and to improve the sustainability of this open-source tool. Another company, Cytomine SA was founded by collaborators to provide services on top of the Cytomine commmunity open-source edition.

I'm very much in favor of basic principles of open science (open access, open data, open source, open hardware) and links to commons (shared resources), e.g. I published this review on Open Practices and Resources for Collaborative Digital Pathology (Frontiers in Medecine, 2019), and we designed BIAFLOWS, a Collaborative Framework to Reproducibly Deploy and Benchmark Bioimage Analysis Workflows (Cell Patterns, 2020; built on top of Cytomine).
 I'm also involved in the BigPicture project (2021-2027, deputy lead of Work Package 4 on "software tools") to establish the biggest open-access database of pathology images to accelerate the development of artificial intelligence in medicine.
I'm also involved in the BigPicture project (2021-2027, deputy lead of Work Package 4 on "software tools") to establish the biggest open-access database of pathology images to accelerate the development of artificial intelligence in medicine.
Software technologies to foster diverse solidarity practices in local communities
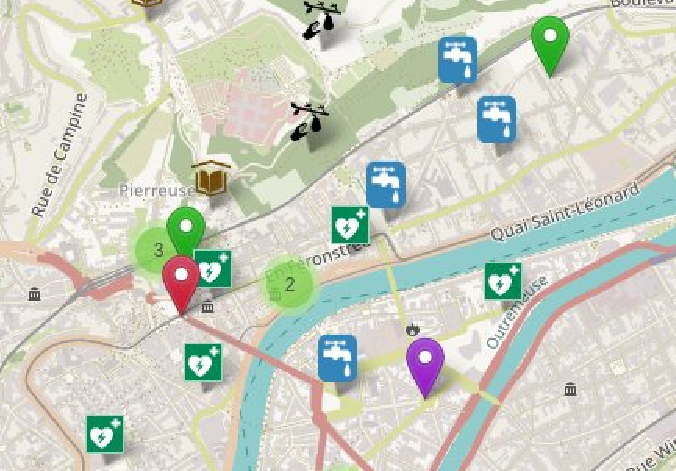
In 2021, I initiated the Shareish (Share and Cherish) project to design and develop an open-source, map-based platform to foster diverse solidarity practices (incl. mutual aid) and strengthen community relations based on human needs rather than market needs, inspired by grassroots movements, principles of gift economy (generalized exchange, indirect reciprocity), and solidarity HCI (human-centered informatics, computer-supported cooperative work). A first paper describes the Shareish platform (Communities & Technologies, 2023), and its source code is available on Github to allow various communities to replicate and adapt the platform to their own needs. While this may seem like a shift in my research interests, I see it as a logical extension of my focus on facilitating collaborative practices, from the scientific community to civil society communities.
Machine/Deep learning for computer vision
Since 2003 we develop general-purpose methods for the recognition of various types of images that share some visual regularities, without relying on too strong assumptions about patterns to recognize and acquisition conditions, and without having to rely on domain experts to design specific features. This book chapter (2013) summarizes our previous work where we combined ensemble of randomized decision trees with random extraction of subwindows (square patches) described by their raw pixel values. To assess versatility of our methods, we often perform large-scale empirical studies e.g. in Pattern Recognition Letters (2016) for image classification, or Nature Scientific Reports (2018) in for interest point (landmark) detection. More recently, we are trying to combine ideas from tree-based methods, deep learning methods, and transfer learning approaches, see e.g. our papers in CVPR-CVMI (2018) and IEEE-JBHI (2020). Since 2016 these algorithms are integrated into the aforementioned CYTOMINE open-source software, and since 2022 we also use deep learning techniques in the Shareish platform to ease user content creation.
Bioinformatics
Between January 2005 and October 2014, I was the GIGA Bioinformatics platform manager. We offered software development and data analysis services to academic (within the GIGA research center and beyond) as well as to industrial researchers. Services included classification of biological/biomedical data (SELDI mass spectra, microarrays, clinical databases, ...) obtained from various medical instrumentation based on machine learning methods. This activity lead to co-authorship of several journal papers (in Journal of Immunology, Proteomics, Annals of the rheumatic diseases, ...) in various application domains (inflammatory diseases, cancer, ...).
Software tools
CYTOMINE is a "Google Maps"-like rich internet application for remote visualization, collaborative and semantic annotation and automated analysis of high-resolution images. It is released under an open-source software license and an official release is maintained by Cytomine cooperative. The software also includes data mining modules encapsulated into Docker containers with our latest algorithm developments for image classification, semantic segmentation, and landmark detection, and a Java module for content-based retrieval. It can be run on big servers for large-scale studies, or on laptops for small-scale works.
BIAFLOWS a collaborative framework to reproducibly deploy and benchmark bioimage analysis workflows (see demonstration server and documentation).
PiXiT was a Java GUI software which implemented our CVPR 2005 image classification method. It is no longer available
and this pyxit reimplementation in Python (command-line) should be used.
Shareish is an open-source, map-based, web platform to foster solidarity, using a combination of spatial databases, Django and Vue.js frameworks. Using Docker technologies, it can be deployed in development mode on personal computers, or production mode on servers.

 orcid.org/0000-0002-9587-1954
orcid.org/0000-0002-9587-1954

 I'm also involved in the
I'm also involved in the 